 Image credit: Unsplash
Image credit: Unsplash
Abstract
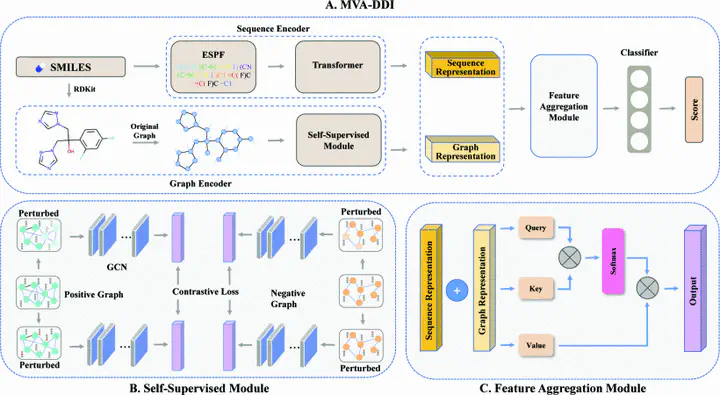
Drug-drug interaction (DDI) plays an increasingly crucial role in drug discovery. Predicting potential DDI is also essential for clinical research. Given the high cost and risk of wet-lab experiments, in-silico DDI prediction is an alternative choice. Recently, deep learning methods have been developed for DDI prediction. However, most of existing methods focus on feature extraction from either molecular SMILES sequences or drug interactive networks, ignoring the valuable complementary information that can be derived from these two views. In this paper, we propose a novel interpretable Multi-View Attention network (MVA-DDI) for DDI prediction. MVA-DDI can effectively extracts drug representations from different perspectives to improve DDI prediction. Specifically, for a given drug, we design a transformer-based encoder and a graph convolutional networkbased encoder to learn sequence and graph representations from SMILES sequence and molecular graph, respectively. To fully exploit the complementary information between the sequence and molecular views, an attention mechanism is further adopted to adaptively aggregate the sequence and graph representations by taking the importance of different views into accounts, generating the final drug representations. Comparison experiments demonstrated that our MVA-DDI 1 model achieved superior performance to state-of-the-art models on DDI prediction.